AMD is excited to announce the release of the AMD ROCm™ 5.7. With AMD ROCm open software platform built for flexibility and performance, the HPC and AI communities can gain access to open compute languages, compilers, libraries and tools designed to accelerate code development and solve the toughest challenges in the world today. The latest version of the AMD ROCm platform adds new functionality while building on your favorite features from ROCm 5.7 and other previous releases. Here we will be introducing HIPTensor and highlighting some of our favorite and newly enhanced features in the rocRAND RNG, and MIGraphX libraries. If you are interested in a more in-depth look at ROCm 5.7, we encourage you to check out the release notes.
HIPTensor
HIPTensor will be first available in ROCm 5.7. HIPTensor is AMD's C++ library for accelerating tensor primitives, powered by the composable kernel backend. Tensor primitives can be used as building blocks in more complex HPC and AI workflows designed to increase their flexibility, shorten development time and improve end-to-end efficiency.
HIPTensor currently supports tensor contraction workflows (bilinear and scale) as well as contextual logging capabilities. This is a first step in providing portability of cuTensor features which can run on AMD GPUs. HIPTensor may generate a contraction plan to solve a given contraction problem, using either sampling or actor-critic tuning methods. This plan may be cached and re-used to solve contraction problems with similar parameters.
HIPTensor currently supports AMD Instinct™ MI100 GPUs with 32-bit floating point data type and AMD Instinct MI200 series GPUs with 32/64-bit floating point data types and up to 4th order tensors. Future development includes adding additional data type support (such as complex floating point, etc.). Further API developments may include additional portability of element-wise operations and reduction workflows, as well as support for importing and exporting cached plans.
Download at GitHub - ROCmSoftwarePlatform/HIPTensor: AMD’s C++ library for accelerating tensor primitives and give it a try.
rocRAND RNG performance increases for discrete distributions
In ROCm 5.7, we’ve introduced a simple but effective optimization for discrete distributions that goes hand-in-hand with recent HIP compiler optimizations that have changed the occupancy of rocRAND kernels:
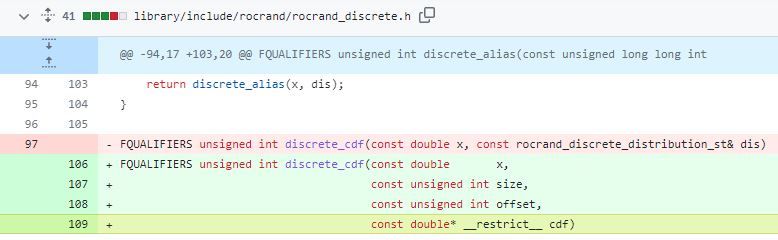
Previously, the entire rocrand_discrete_distribution struct was passed by reference to the discrete table function (discrete_cdf). However, with recent compiler changes, this resulted in suboptimal kernel occupancy. The solution was to pass only the members of the rocrand_discrete_distribution struct that was needed by the kernel, and more importantly, to prefix the cumulative distribution function pointer (cdf) with the __restrict__ keyword.
The __restrict__ keyword in C indicates to the compiler that for the lifetime of the restricted pointer, no other pointer will be used to access the data that the restricted pointer points to. This allows the compiler to make new optimizations that previously would not have been possible.
MIGraphX Introduction
MIGraphX is a graph compiler focused on accelerating machine learning inference for AMD hardware. MIGraphX accelerates machine learning models by leveraging several graph-level transformations and optimizations. Internally, MIGraphX first parses ONNX or TensorFlow models into an internal graph representation (IGR). It then performs optimizations on the IGR to improve the model’s inference performance. Some of the optimizations include :
- Operator fusion
- Arithmetic simplifications
- Dead-code elimination
- Common subexpression elimination (CSE)
- Constant propagation
After performing these transformations, MIGraphX produces code for the AMD GPU either by calling other ROCm libraries, such as MIOpen or rocBLAS, or by creating HIP kernels for a particular operation. MIGraphX can also target CPUs by using the DNNL or ZenDNN libraries.
MIGraphX provides easy-to-use APIs in C++ and Python to import machine models in ONNX or TensorFlow. Users can compile, save, load, and run these models using MIGraphX’s C++ and Python APIs.
New features
Dynamic batch: Oftentimes one might want to run a model with multiple batch sizes depending on the data input. MIGraphX provides a new feature, dynamic batch, that allows users to input data of multiple different batch sizes into a MIGraphX compiled model. To use the feature, a user must provide a range of batch sizes to optimize for during model compilation.
Try it for Yourself
See for yourself the power of ROCm 5.7. Download the latest version.
Contributors
Chris Austen
Charlie Lin
Steven Zhang
Stanley Tsang